
The DECIPHER Project
@deciphergenomic
ID: 4501798349
https://www.deciphergenomics.org 16-12-2015 10:11:03
414 Tweet
2,2K Takipçi
108 Takip Edilen

.The DECIPHER Project v11.25 is here. This update introduces new features including the display of functional data from multiplexed assays of variant effects (MAVEs) to assist variant interpretation and much more. ebi.ac.uk/about/news/upd…




Unique very own Sarah Wynn talking about the impact of how we name rare genetic conditions on families at ClinGen and The DECIPHER Project Curating the Clinical Genome Conference in Baltimore. #ccg2024 Genetic Alliance UK Breaking Down Barriers Bill Newman


Jeremy Arbesfeld presenting pivotal work ccg2024g about a framework for mapping MAVE data, which has allowed DECIPHER and other resources to display MaveDB data ClinGen Atlas of Variant Effects Alliance Alan Rubin





A Genome Aggregation Database CNV genome browser track is now available which displays rare coding autosomal CNVs called from exome sequencing of 464,297 individuals included in gnomAD v4.1. pic.x.com/7ftaxkmy0l

Our gene pages now link out to Open Targets Platform associations pages — view diseases associated with your gene and explore relevant genetic, pathway, drug, bibliography evidence and more. pic.x.com/uu4okzvsxu



Girish Katta Zane Lombard The DECIPHER Project Aimé Lumaka African Rare Diseases Initiative H3ABioNet Victoria Nembaware AfSHGenetics AfricanSocHumGenet ACMG Watch the trailer to discover more about #FLGenomicVariantsDiversity from our educators! Hear about what you will learn, live over 3 weeks, through hands-on exercises delivered via FutureLearn. 🎥 i.mtr.cool/stfhxputzj


UTRannotator scores are displayed on annotation tabs for variants in 5’untranslated regions that create/disrupt upstream open reading frames Nicky Whiffin Xiaolei Zhang 张小蕾 #noncoding
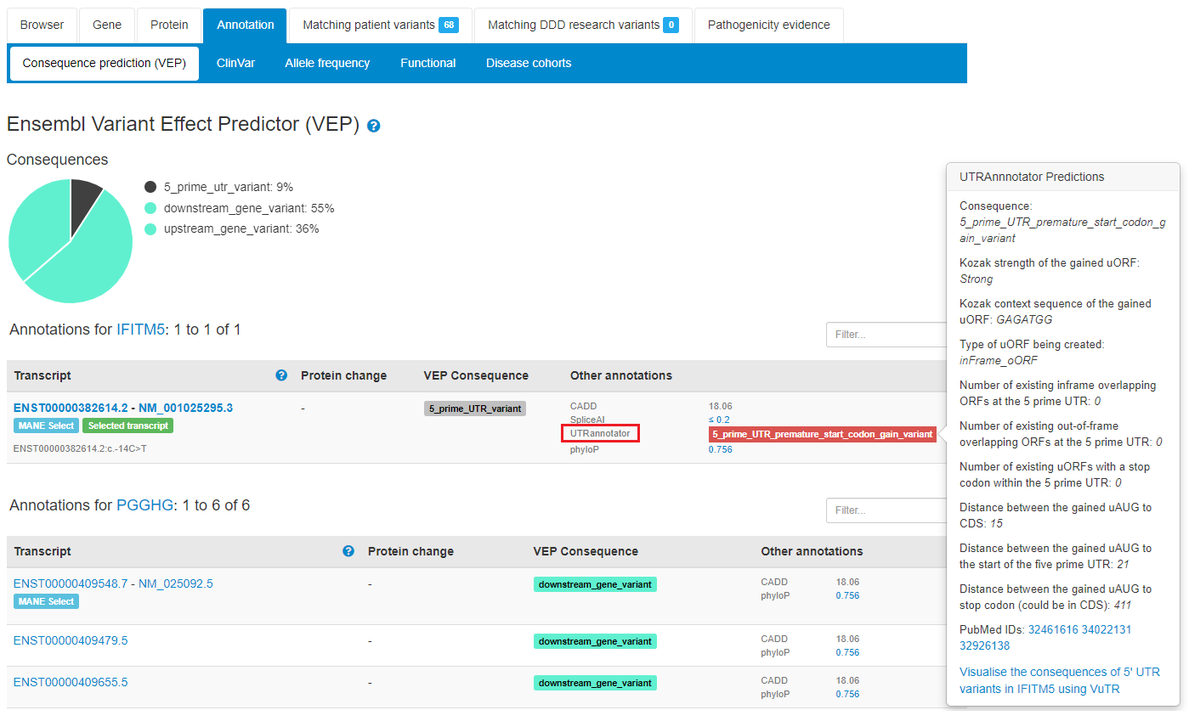

The structural variant genome track has been updated to Genome Aggregation Database v4.1 – 1,199,117 high quality structural variants identified in 63,046 genomes from unrelated individuals


Regional missense constraint data displayed in the genome and protein browser tracks has been updated to Genome Aggregation Database v2.1.1



UK Clinical Genetics BSGM UK Cancer Genetics Group The DECIPHER Project Helen Firth James Fasham Kaitlin Samocha AGNC Nicky Whiffin Diana Baralle Why attend #ClinicalGen25? 🗣️ We'll be joined by leading speakers including: Caroline Wright, Clare Turnbull- Prof/NHS Dr, James Fasham, and James Ware 🖥️ Master online tools (The DECIPHER Project) 🧬 Workshops on cancer and cardiac genetics using DECIPHER 🗓️ Join us 3-5 February 2025






