Xiang Zhou
@xiangzhoulab
Genetics, Genomics, Statistics
ID: 233160369
http://xiangzhou.github.io 02-01-2011 13:22:40
33 Tweet
657 Followers
149 Following

Our latest with Mengjie Chen on imputing single cell RNAseq data while preserving single cell level expression variability: tinyurl.com/yah9gvcs Genome Biology



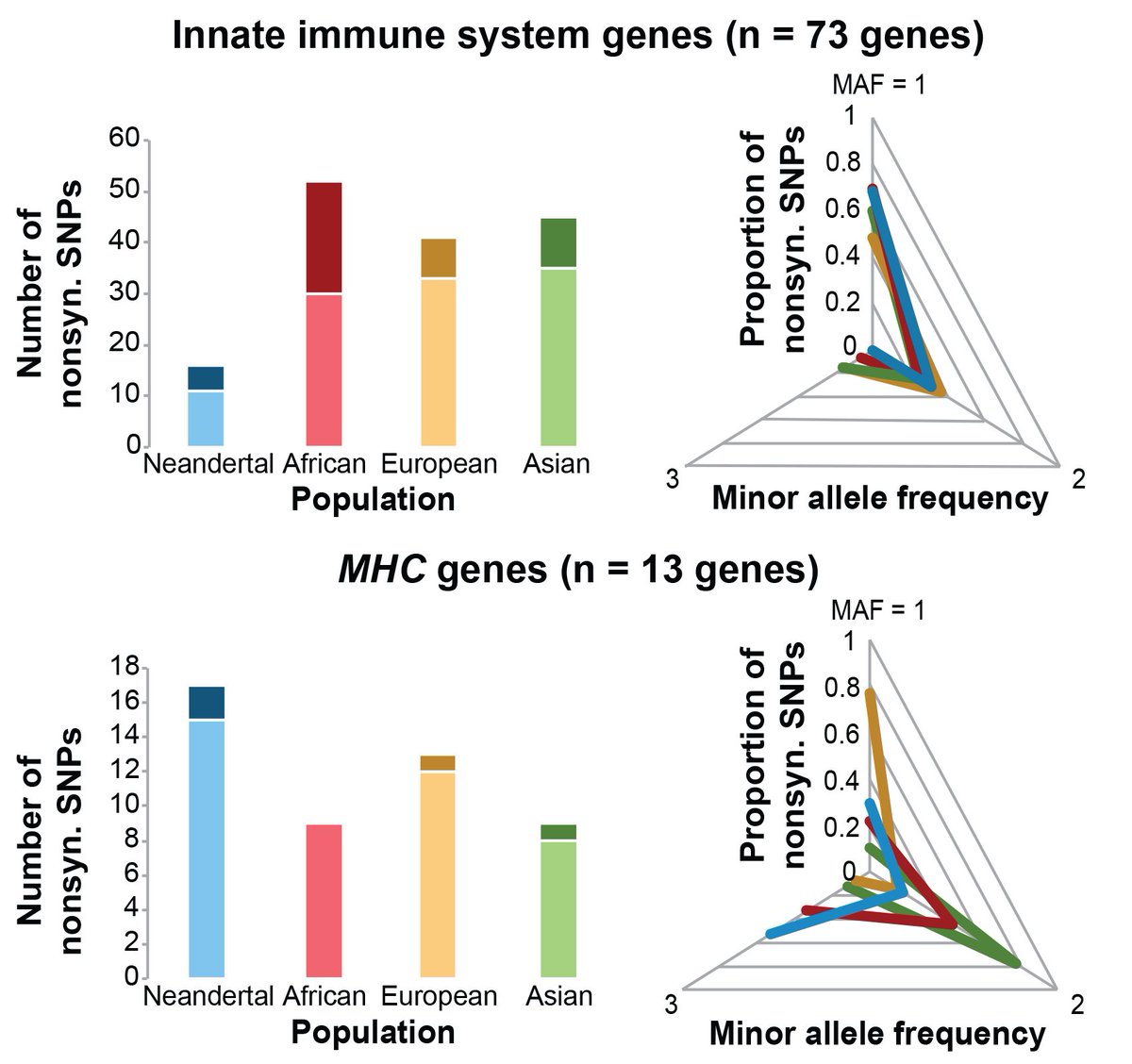
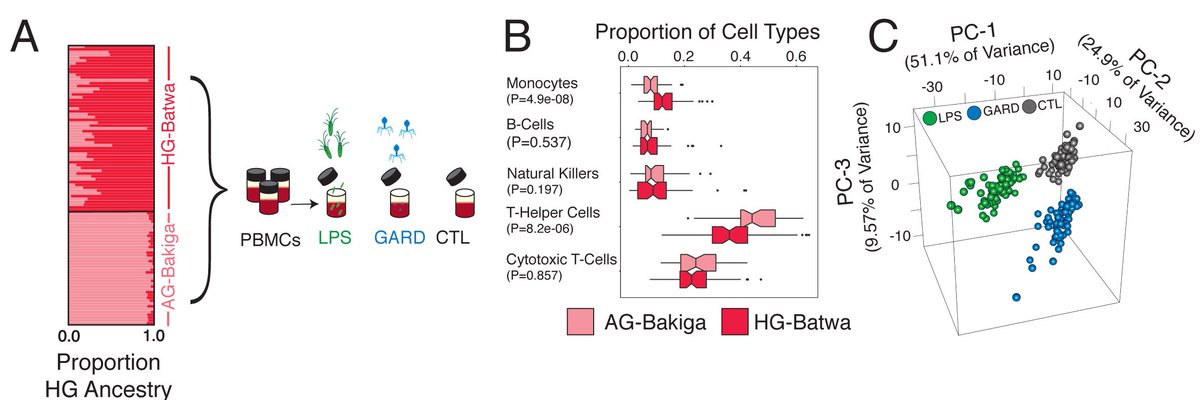
Excited to share our most recent work led by @GeneScientist17: Natural selection contributed to immunological differences between human hunter-gatherers and agriculturalists. Great collab with George (PJ) Perry lab with contributions from many others biorxiv.org/cgi/content/sh… 1/5


Exciting collaboration with Jenny Tung on a computational method for integrative allelic specific methylation analysis and mQTL mapping, with applications to baboons and wolves genomebiology.biomedcentral.com/articles/10.11…

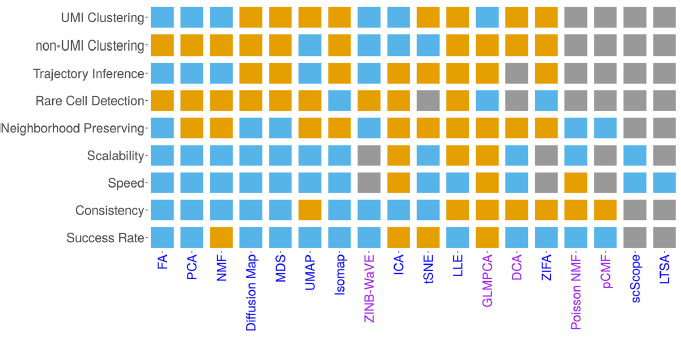
Exciting work lead by Shiquan Sun, PhD; we were also quite surprised to see the results.

A benchmarking of dimensionality reduction techniques for scRNA-seq data from Shiquan Sun, PhD, @xzlab_org and co. They compare 18 methods on 2 simulated and 30 real datasets. There is not one method that works well across all tasks. genomebiology.biomedcentral.com/articles/10.11…


Very excited on this work with Mengjie Chen Mengjie Chen

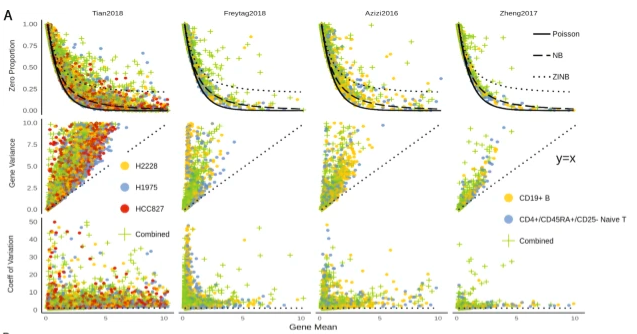
HIPPO, a heterogeneity-inspired preprocessing tool for analyzing RNA-seq data with UMIs, from Kim, @xzlab_org, and Mengjie Chen. Drop-outs follow a Poisson distribution, and can be used to measure cell-type heterogeneity, before normalization. genomebiology.biomedcentral.com/articles/10.11…




Congrats Helena Kilpinen ([email protected]) Oliver Stegle Daniel Gaffney and others! biorxiv.org/content/early/…



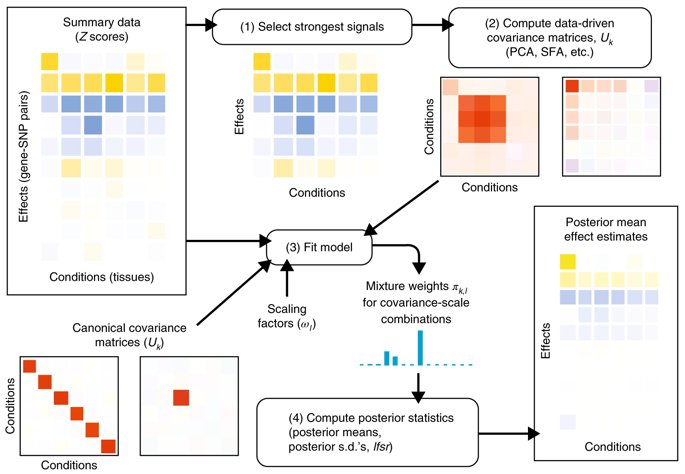
Our paper on the MArginal ePIstasis Test. Thanks for helping to get this out, PLOS Genetics! journals.plos.org/plosgenetics/a…