
RGB Lab @ MIT
@RGBLabMIT
Official Twitter for the Gómez-Bombarelli group @MIT_DMSE | We use atomistic simulations and ML for accelerated materials design | Managed by group members
ID:1579875531339706368
http://gomezbombarelli.mit.edu/ 11-10-2022 16:44:47
124 Tweets
1,4K Followers
142 Following
Follow People



I feel personally attacked. 😅
Seriously though - sorry Ben Blaiszik, we’re just not that good at planning ahead.


📢 New preprint! Our latest work enables 'alchemy' - ∂[energy]/∂[element] in ML potentials like MACE. We model solid solutions and conduct alchemical free energy simulations. arxiv.org/abs/2404.10746
RGB Lab @ MIT #compchem #machinelearning
![Juno Nam (@junonam_) on Twitter photo 2024-04-30 18:11:33 📢 New preprint! Our latest work enables 'alchemy' - ∂[energy]/∂[element] in ML potentials like MACE. We model solid solutions and conduct alchemical free energy simulations. arxiv.org/abs/2404.10746 @RGBLabMIT #compchem #machinelearning 📢 New preprint! Our latest work enables 'alchemy' - ∂[energy]/∂[element] in ML potentials like MACE. We model solid solutions and conduct alchemical free energy simulations. arxiv.org/abs/2404.10746 @RGBLabMIT #compchem #machinelearning](https://pbs.twimg.com/media/GMbmdKebIAAep2_.jpg)

Check out our recent work on understanding and predicting atomic orderings in multicomponent oxides just published at Cell Reports Physical Science! Stay tuned—we will share a very cool follow-up study soon. RGB Lab @ MIT


Design of cytotoxic T cell epitopes for enhancing antigen processing and presentation, enabled by machine learning of human degrons and cytosolic delivery with two anthrax proteins
NEW #ASAP by PenteluteLab RGB Lab @ MIT & team
Read it here: go.acs.org/93q


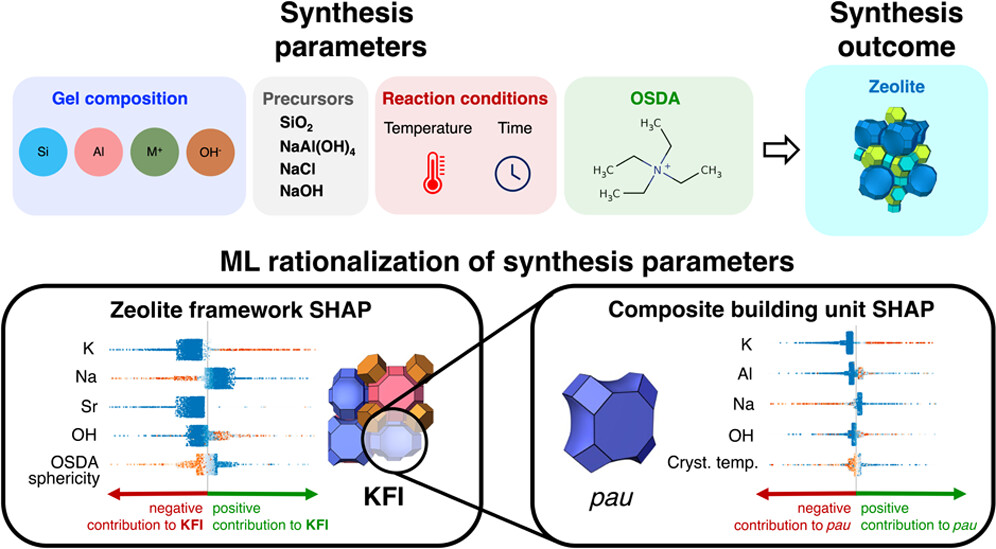
A Large Open-Source Zeolite Synthesis Dataset Enables Machine Learning Rationalization of Zeolite Structure-Synthesis Relationships
NEW #ASAP by DMSE at MIT Olivetti Group @ MIT RGB Lab @ MIT MIT ChemE Dept Román Lab @ MIT ITQ (UPV-CSIC)
Read it here: go.acs.org/92l

Registration is now open for our 5-day Simons workshop taking place in two months (June 10th-14th) in Berkeley (w Aditi Krishnapriyan ).
AI≡Science: Strengthening the Bond Between the Sciences and Artificial Intelligence simons.berkeley.edu/workshops/aisc…


Great single-author paper by Michael Skinnider. We've suspected that bad SMILES were a 'canary in the coal mine' for lack of reliability in downstream tasks. It is great to see quantitative work about it. Being able to be wrong actually helps the models be right.

GenAI lowers the barriers to designing new chemicals. Akshay Subramanian, Wenhao Gao, Regina Barzilay, Jeff Grossman, Tommi Jaakkola, Stefanie Jegelka, Mingda Li, Ju Li, Wojciech Matusik, Elsa Olivetti, Connor W. Coley & RGB Lab @ MIT discuss opportunities and risks: mit-genai.pubpub.org/pub/681kpeoa/r…



In the talk from RGB Lab @ MIT we will learn how to 'differentiate everything everywhere all at once' and how we can use that to obtain the best neural networks possible #virapid2024



Rui Tan's excellent work shows that we currently lack a fully reliable uncertainty quantification method for MLIPs, as no single method manages to outperform the others.
doi.org/10.1038/s41524…
RGB Lab @ MIT npj Journals

I'm happy to share that our paper 'Autonomous, multiproperty-driven molecular discovery: From predictions to measurements and back' was published yesterday in Science Magazine: science.org/doi/10.1126/sc…

Thanks for the shoutout Mie Andersen ! 🤩
Nature Computational Science
nature.com/articles/s4358…

Researchers in RGB Lab @ MIT devised a machine-learning method to investigate how materials behave at their surfaces. The approach could help in developing compounds or alloys for use as catalysts, semiconductors, or battery components. buff.ly/4a9chKS


📢RGB Lab @ MIT and colleagues from DMSE at MIT propose a Markov-chain method to predict surface phase diagrams of multi-component materials, enabling fast and accurate sampling of material surfaces. nature.com/articles/s4358…
👉rdcu.be/ds1rR







