WeiniHuang
@huang_weini
PI in mathematical biology, cancer evolution, theoretical biology, species coevolution, polymorphism under random mutations
ID: 3232028201
https://scholar.google.com/citations?hl=en&user=T9RvGuoAAAAJ&view_op=list_works&sortby=pubdate 03-05-2015 22:27:20
706 Tweet
1,1K Followers
730 Following

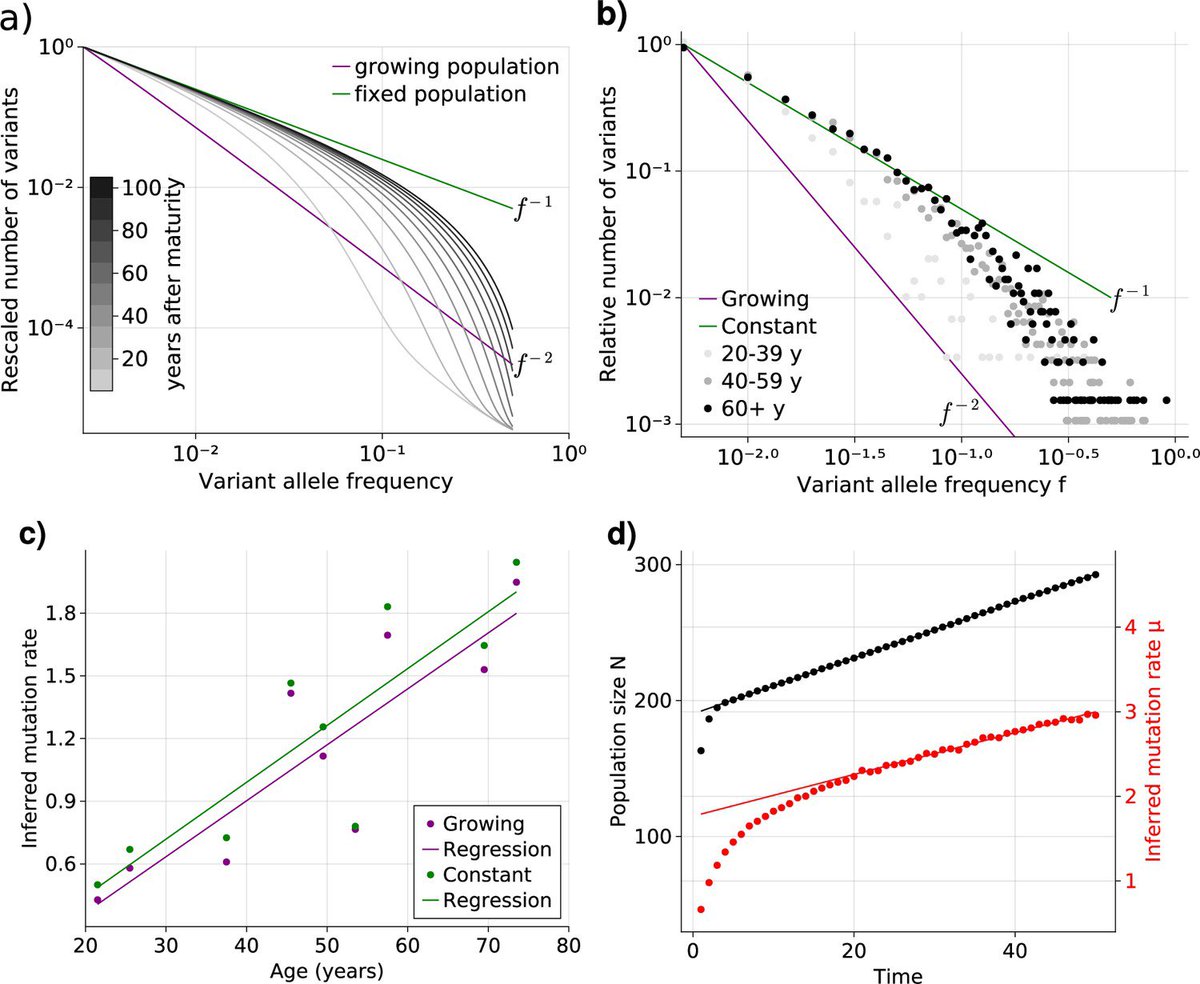
Thanks Arda Durmaz and Valeria Visconte for a nice summary highlighting our theory integrating bulk and single cell data. elifesciences.org/articles/95513 eLife - the journal work done together with Benjamin Werner , Nathaniel Mon Pere and Marius Moeller

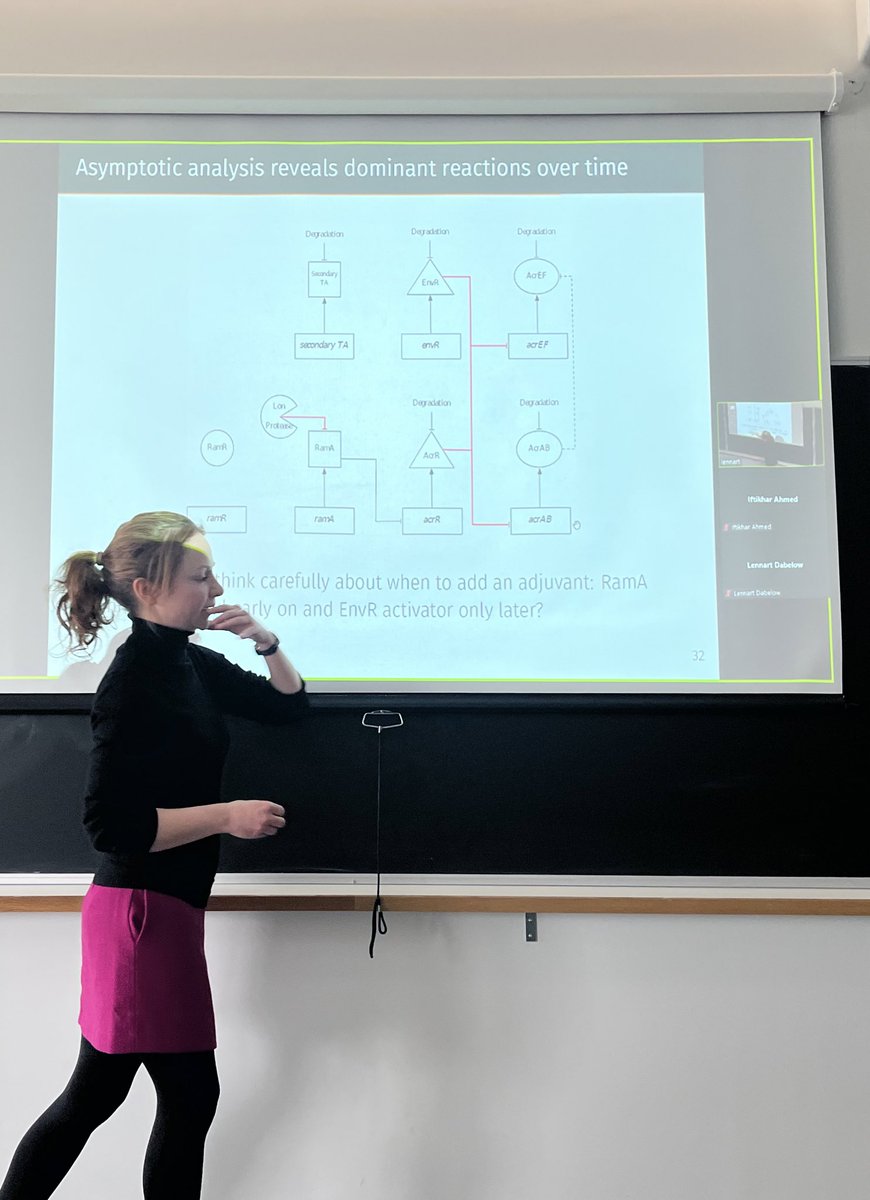
great to have Sara (Jabbari) School of Mathematics coming to qmul talking about using mathematical modelling to aid in the development of anti-resistance treatments, applying drugs in a more reasonable order under immune response and finding a better target in efflux pump network.


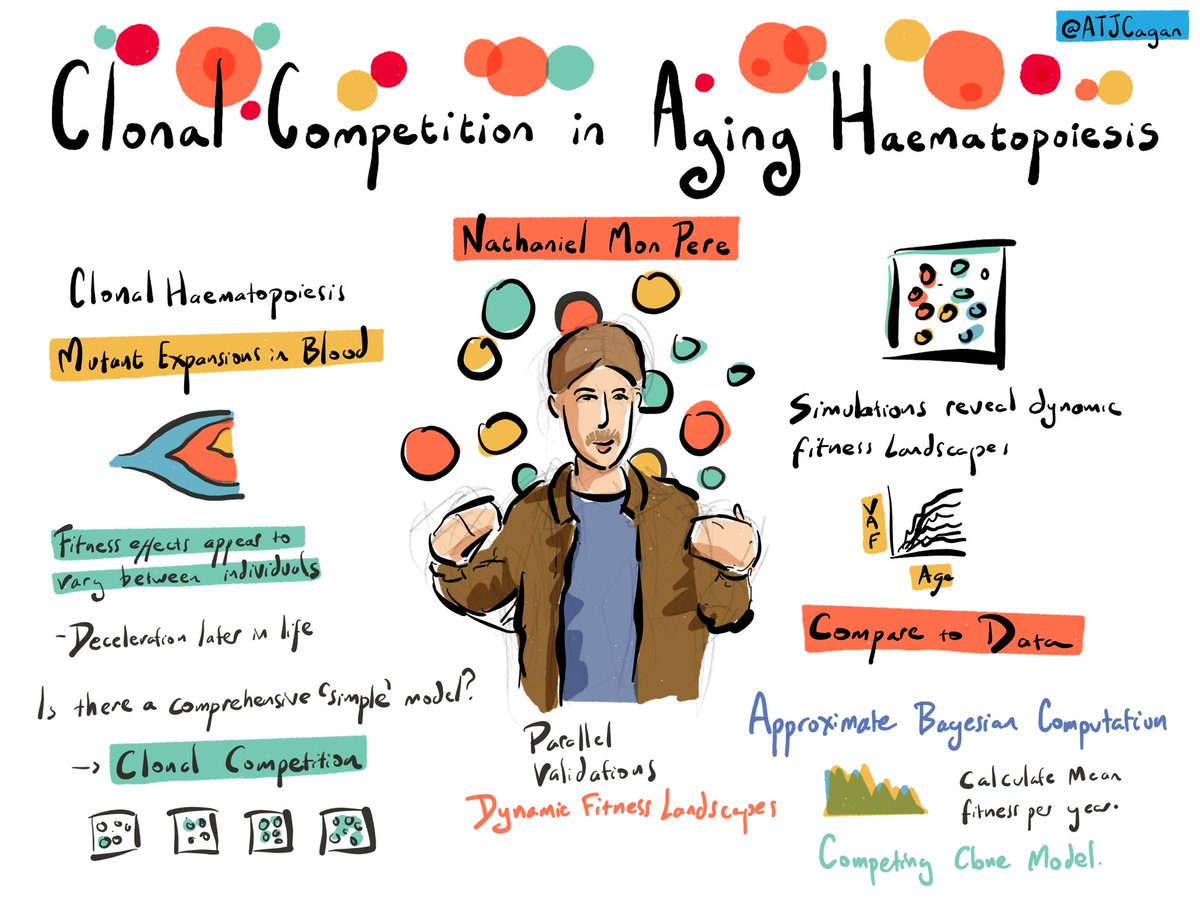
I am very excited about our newest preprint on the emerging dynamic fitness landscape of haematopoietic stem cells trough clonal competition. biorxiv.org/content/10.110… This work has been pushed by the fantastic Nathaniel and Francesco Terenzi.

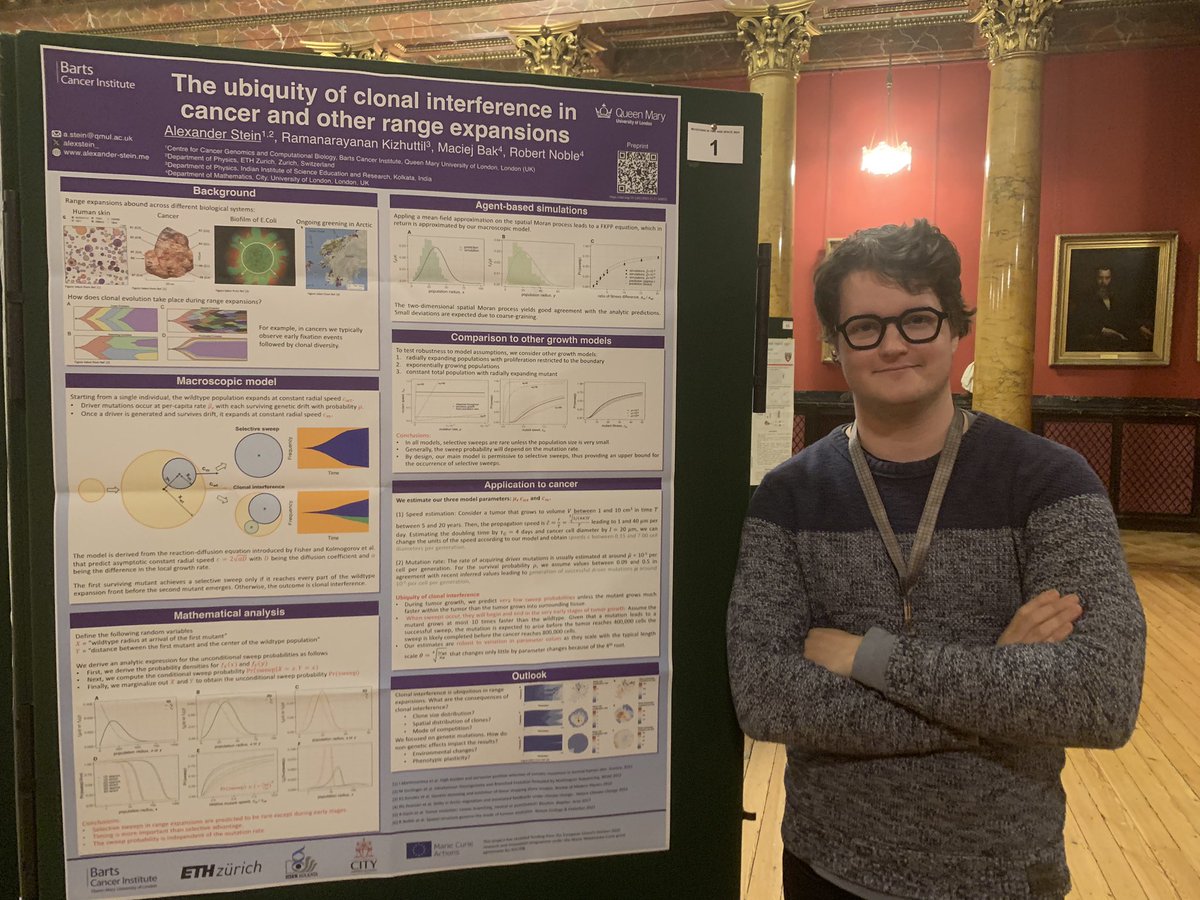
Today Nathaniel will also talk about this work during the very first session of the mutationmeeting and Francesco Terenzi will have a poster later today in the afternoon presenting more details on the SFS modelling.



Parasites: the silent puppeteers of the food web? Eizaguirre Lab and Dr WeiniHuang shed light on how these tiny organisms influence predator-prey dynamics in their new study. royalsocietypublishing.org/doi/10.1098/rs…


I am very happy to be part of the team and continue to learn about the evolution of clonal organisms . Its also great to see that methods of "classical" species evolution and somatic (cancer) evolution are of mutual benefit Barts and The London, Queen Mary Cancer Research UK UK Research and Innovation Queen Mary University of London Cancer Grand Challenges