Arang Rhie
@ArangRhie
Staff Scientist in Genome Informatics
ID:817382322274070528
06-01-2017 14:48:26
490 Tweets
1,2K Followers
153 Following
Follow People

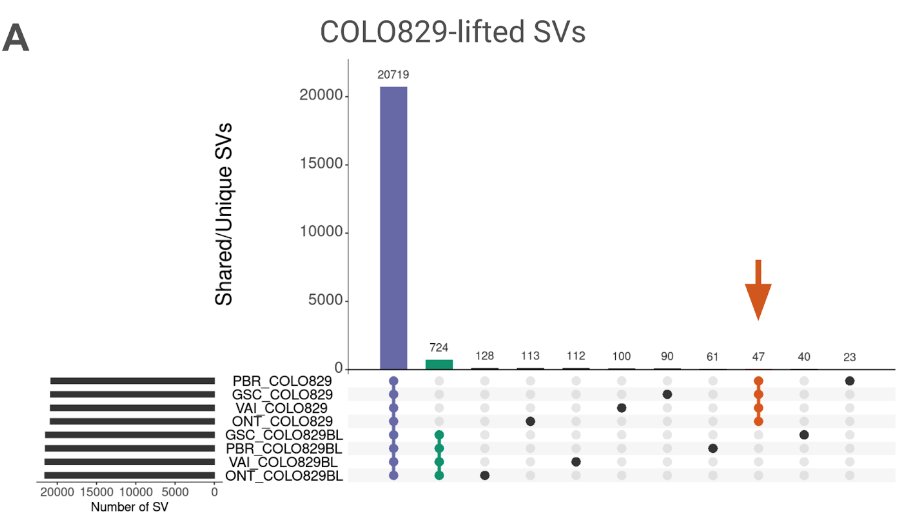
New work on #Cancer variant prioritization using #T2T . It helps & we can overcome variant annotation issues! + An updated somatic SV benchmark for COLO829/COLO829BL across 4 replicates Grch38 +T2T!
New preprint: medrxiv.org/content/10.110…
Luis Paulin, PhD BCM HGSC Kieran O'Neill etc!

Preprint on Exploring gene content with pangenome gene graphs: arxiv.org/abs/2402.16185. It describes pangene for building gene graphs and for calling gene-level variations which can be found at pangene.bioinweb.org.
Pleasant collaboration with Maximillian Marin and Maha Farhat.

Memphis Pangenomics days, 2024 edition! Pangenome Workshop (May 18-19), Conference (May 20), & Biohackathon (May 21-22). pangenome.github.io/MemPanG24/ #Pangenomics #Bioinformatics #MemPanG24




1/5 We (Nathaniel Brown, Omar Ahmed, Travis Gagie, and Ben Langmead) developed Movi, a cache-efficient full-text pangenome index. It's the fastest full-text index for pangenomes, particularly appropriate for adaptive sampling where time budget is important.

I am excited to share that National Human Genome Research Institute has created a new publicly available digital archive and search aid for accessing documents related to the history of genomics!

So happy to finally see our MAGE paper out on bioRxiv!! I'm so excited about this data set; I hope this will be a valuable resource for the field of functional genomics. Congrats as well to my co-authors Surya Chhetri, Michael Tassia, Arjun Biddanda, Alexis Battle, and Rajiv McCoy


Just in time following an incredible #ASHG23 , we're excited to share our work on quantifying sources of gene expression variation across a global human cohort! At the helm of this project is Dylan Taylor in Rajiv McCoy's lab!
biorxiv.org/content/10.110…

Genome Research is now encouraging submissions for a Special Issue on Long-read DNA Sequencing Applications in Biology and Medicine, Guest Edited by AnaConesa, Alexander Hoischen, and Fritz Sedlazeck bit.ly/grcallforpapers #longreads #T2T #LongTREC #Hitseq #ASHG23





Our preprint is out! We hacked the Oxford Nanopore sequencer to read amino acids and PTMs along protein strands. This opens up the possibility for barcode sequencing at the protein level for highly multiplexed assays, PTM monitoring, and protein identification!
biorxiv.org/content/10.110…

It was wonderful celebrating #sammies2023 honorees last evening at the Kennedy Center in DC! NHGRI was particularly proud of Adam Phillippy, Sergey Koren, and Arang Rhie for being Sammies finalists for their groundbreaking contributions to the Telomere-to-Telomere Consortium.


I am thrilled to share our study on the de novo assembly and characterisation of 43 diverse human Y chromosomes, published today in nature, together with the first #T2T Y assembly from Adam Phillippy, Arang Rhie and co-authors nature.com/articles/s4158…

.@Genome_gov researchers have assembled the first, fully complete human #YChromosome sequence! This sequence reveals genetic factors with implications for fertility. genome.gov/news/Y-chromos… (Thread)


Our latest methods paper 'Minmers are a generalization of minimizers that enable unbiased local Jaccard estimation' (aka MashMap3) is out with Bryce Kille Erik Garrison Todd J Treangen! There's an interesting backstory here 🧵 \ biorxiv.org/content/10.110…