James Ferguson
@Psy_Fer_
Bioinformatician/Genomics Software Engineer @GarvanInstitute, Views my own.
Mastodon @[email protected], https://t.co/K5efuEeBwt
ID:761420960046080000
http://jmferguson.science 05-08-2016 04:37:58
13,2K Tweets
3,9K Followers
1,2K Following
Follow People



Genome assembly in the telomere-to-telomere era go.nature.com/4aLqIED #Review by Heng Li Heng Li & Richard Durbin Richard Durbin
Dana-Farber DBMI at Harvard Med Cambridge University

Our review paper on 'Genome assembly in the telomere-to-telomere era' published online in Nature Reviews Genetics. It amazes me how much assembly has progressed in the past four years – often ~100X improvement in contiguity and base accuracy and with haplotypes and hard regions resolved!

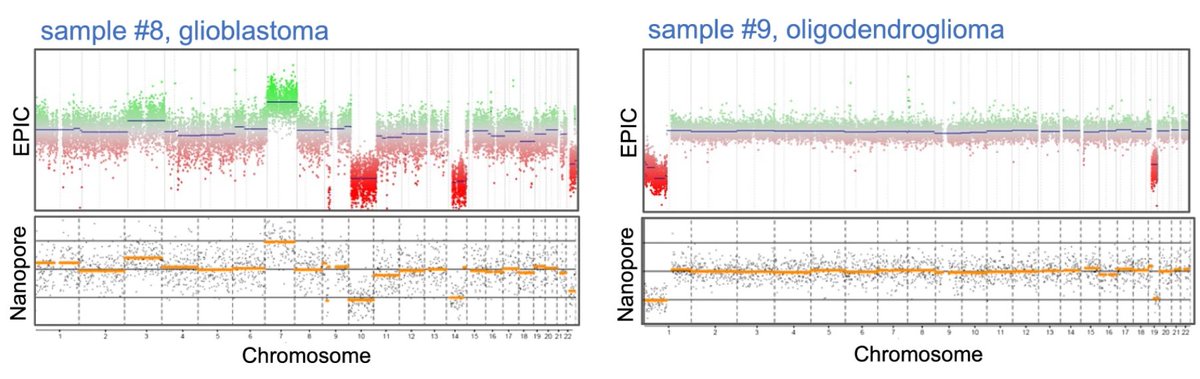
Always thought #Nanopore sequencing from #FFPE is impossible? Explore our paper: tinyurl.com/ffpe-nanopore. Methylation & CNV profiling of CNS tumors. Great collaboration with Ulrich Schüller and Leo. Oxford Nanopore

Working at UNSW & surrounds? Want to meet others working in #Bioinformatics ? Help is at hand! Join us this coming May 9th,2024 for some ☕️🍵🍪 and 💻🧬
RSVP here: forms.gle/zw3hBc7aLGaFiT…
Sponsored by UNSW Computing, jointly organised with the #RegRNALab UNSW Biotechnology & Biomolecular Sciences. Please share!



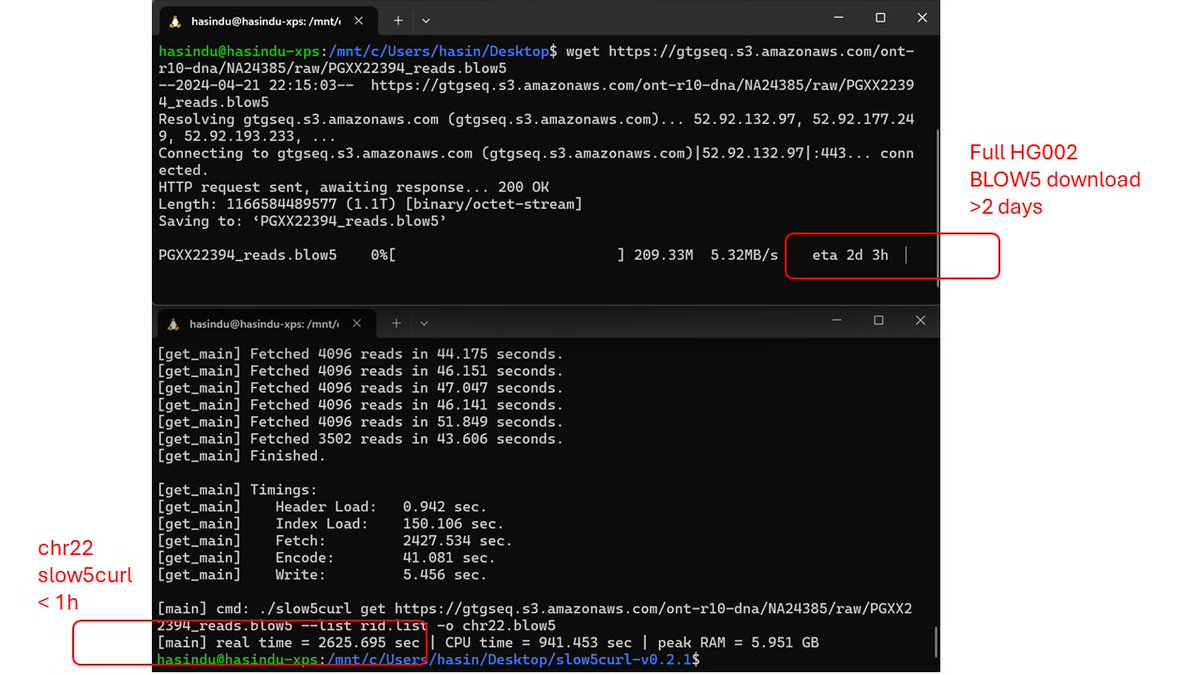
Copy+paste following commands to get chr22 from a 1TB BLOW5. <1h on a ~50Mbps.
wget github.com/BonsonW/slow5c… -O s5curl.tgz
tar xf s5curl.tgz && cd slow5curl-v0.2.1
wget raw.githubusercontent.com/BonsonW/slow5c… -O rid.list
./slow5curl get gtgseq.s3.amazonaws.com/ont-r10-dna/NA… --list rid.list -o chr22.blow5


Come and meet stellar cast of international and national speakers at #AGTA24 Australasian Genomic Technologies Association AGTA
Register 👉agta.arinex.one/pages/registra…
Thanks Ramaciotti Centre for Genomics Martin Smith for the PR
#genomictechnologies
ABACBS folks - come & join us. 10% discount! T&Cs apply.

Ready for >Q30 ONT reads? Herro error correction now supports both R9.4.1 and R10.4.1. simplex reads w/ Dominik Stanojevic Dehui Lin (CHM13 -R9.4.1) #nanopore #longreads


Demuxafy: improvement in droplet assignment by integrating multiple single-cell demultiplexing and doublet detection methods.
url.au.m.mimecastprotect.com/s/dmyICmO5nJhp…
Led by Drew Neavin in collab with the sc-eQTLGen Consortium Lude Franke Shyam Prabhakar Lab Jimmie Ye Davis McCarthy Marta Melé Martin Hemberg

Happy to share our new article on using Oxford Nanopore adaptive sampling to profile DNA methylation markers! doi.org/10.1016/j.fsig… Special thanks to Eduardo Eyras, Cameron Jack, Runa Daniel, Dennis McNevin, and other collaborators at John Curtin School of Medical Research, ANU #DNAmethylation #Bioinformatics

Our #slow5curl paper is out! academic.oup.com/gigascience/ar…
Big projects storing hundreds/thousands of Oxford Nanopore signal datasets on cloud storage like Amazon Web Services #s3 will be able to save 💸,⏲️ and bandwidth while improving the accessibility of datasets to those with limited computing ..










