Ilia Igashov
@igashov
PhD student at LPDI, EPFL 🇨🇭 Artificial Intelligence & Structural Biology
ID:1299969438356320256
http://igashov.github.io 30-08-2020 07:17:38
118 Tweets
884 Followers
290 Following



Very excited to present SynFlowNet next week at GEMBio Workshop #ICLR2024 ! We tackle the synthesisability issue in molecular design by training a GFlowNet with an action space of chemical reactions. Thanks to my coauthors Charlie Harris Julien Roy Emmanuel Bengio and Pietro Lio (1/n)

Our method for molecular linker design with diffusion models (DiffLinker) got published in Nature Machine Intelligence 🥳
nature.com/articles/s4225…
Thanks a lot to our amazing coauthors Hannes Stärk Clément Vignac Arne Schneuing Víctor Garcia Satorras Pascal Frossard Max Welling Michael Bronstein Bruno Correia

Professor Michael Bronstein (Michael Bronstein @ICLR in Vienna) has co-authored the article 'Equivariant 3D-conditional diffusion model for molecular linker design' published today in Nature Machine Intelligence (@NatMachIntell).
Read here: nature.com/articles/s4225…
#compscioxford nature

A packed room and exciting talks - the 4th Applied Machine Learning Days AI & the Molecular World track was a full success!
Many thanks to the excellent speakers, to the participants, and to Rebecca Neeser Ilia Igashov @ ICLR 2024 🇦🇹 Arne Schneuing @ ICLR 2024 Damiano Sgarbossa @ICLR2024 and Bruno Correia for the organisation! #AMLD2024EPFL
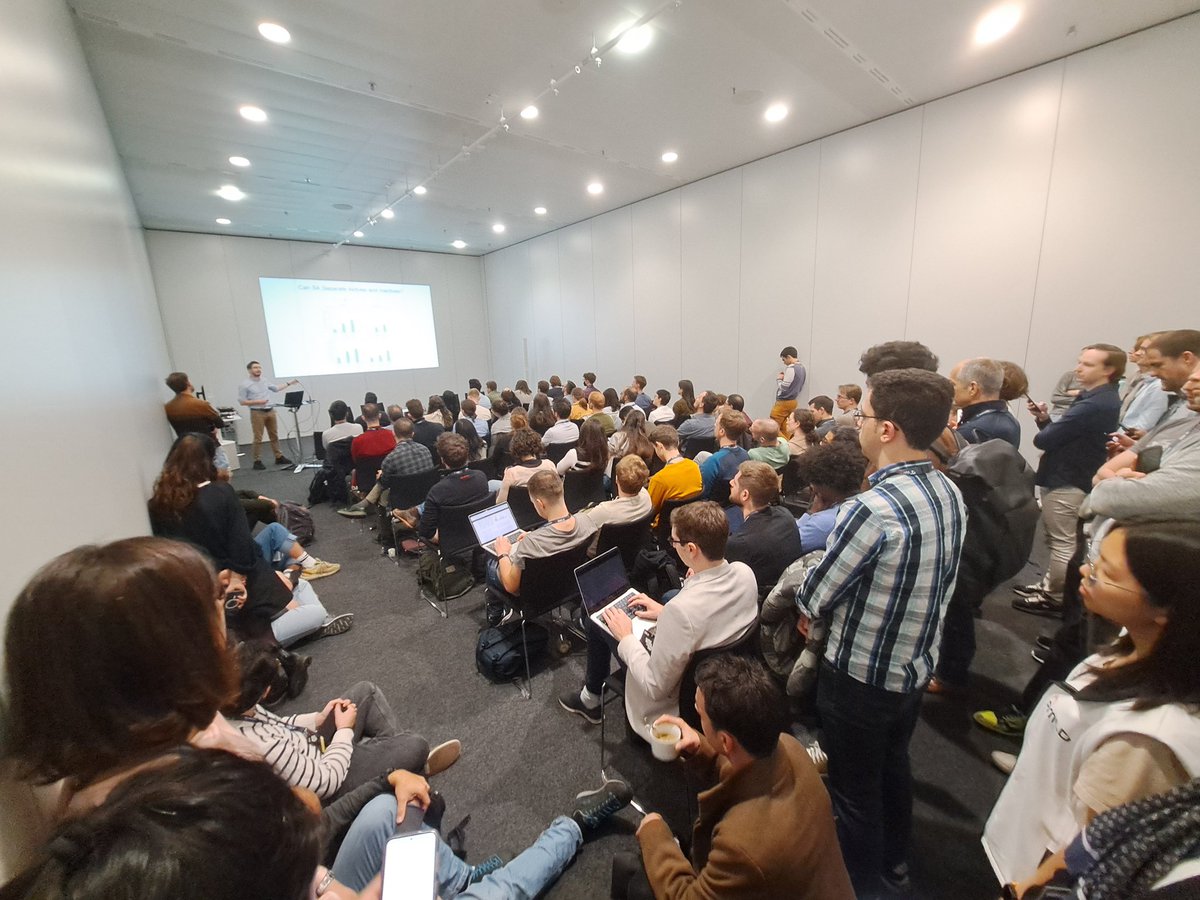


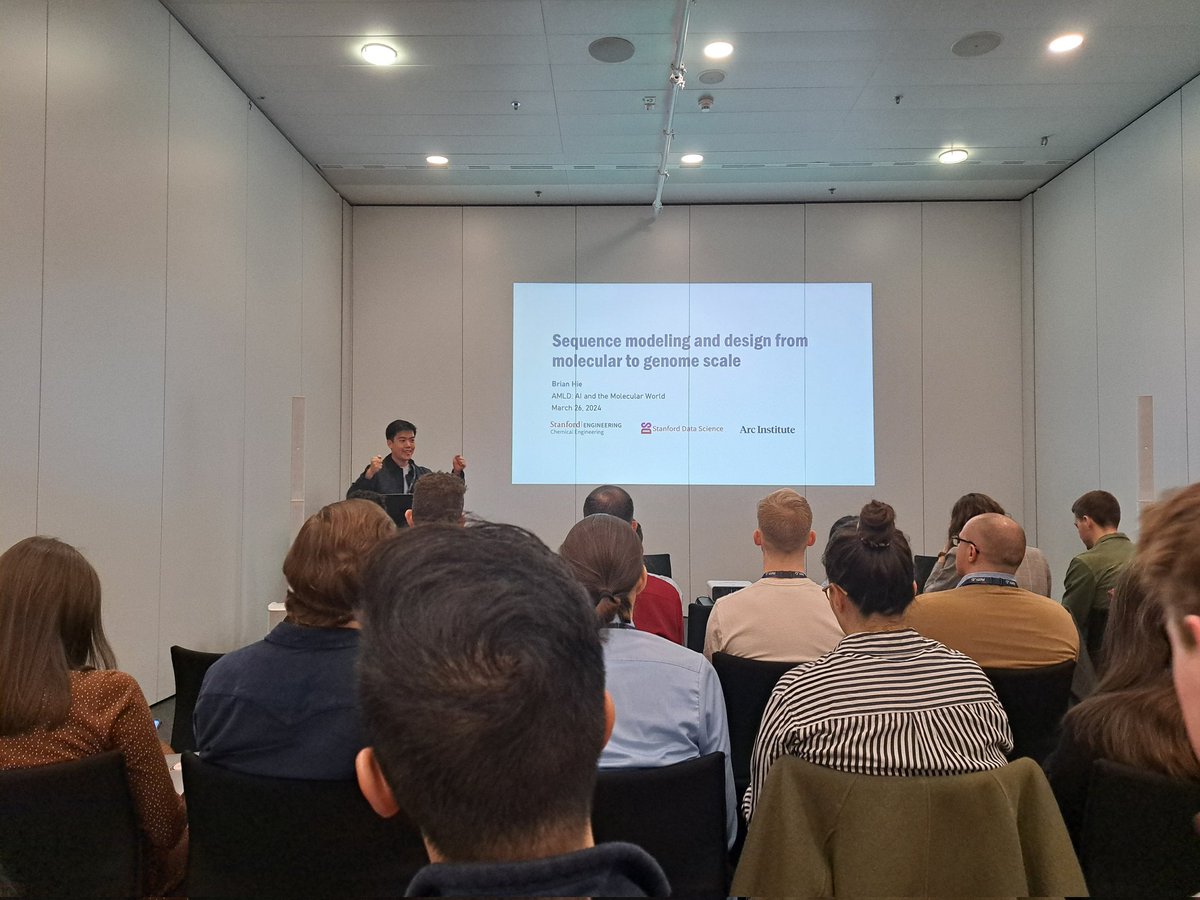
The #AMLDEPFL2024 AI & Molecular World track has started!
After Rebecca Neeser's introduction, Brian Hie gave a fascinating talk on Evo - a genome-scale LLM!
Applied Machine Learning Days


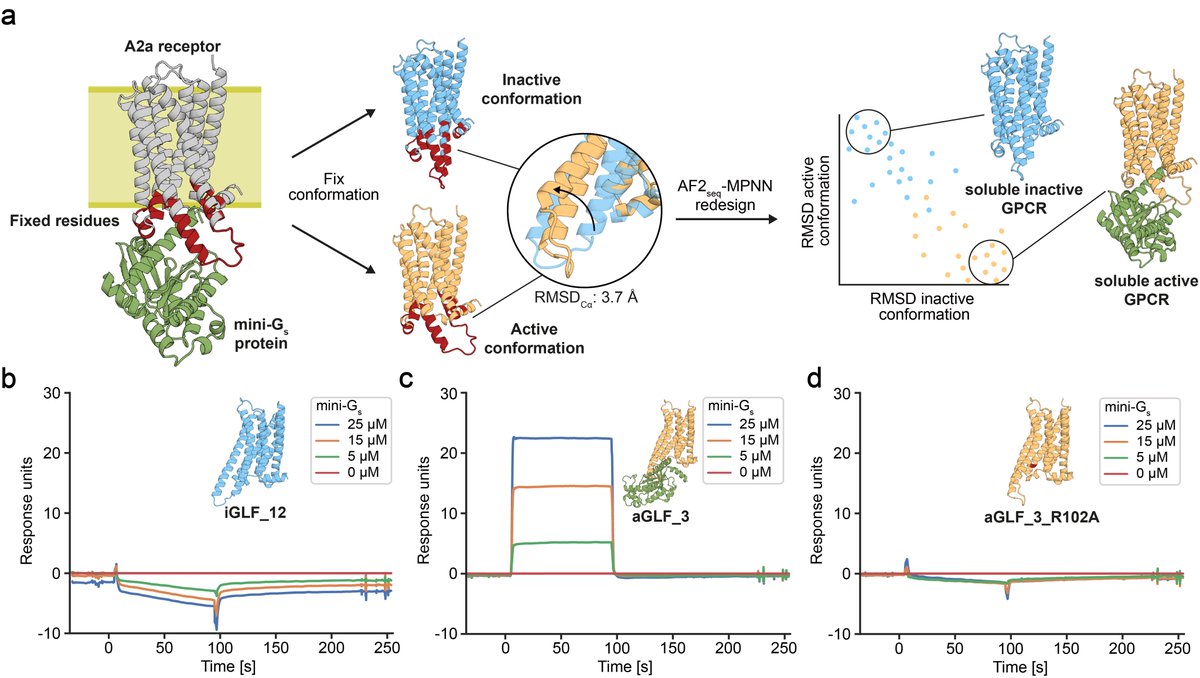
THEY SAID WE COULDN'T DO IT........well, no one actually said that. But in our updated manuscript with Casper Goverde, Nico Goldbach, and Bruno Correia we show you can now design soluble analogues of membrane proteins with preserved functional features!
biorxiv.org/content/10.110…

Hi #GraphLearning folks! I am looking for a node-level autoencoder for molecular graphs. With a possibly simple decoder like in VGAEs (Kipf & Welling, 2016). I guess there should be something widely used for molecules but I am struggling to find a good reference. Any ideas?


We’ll be hosting the 'AI & the Molecular World' track at #AMLDEPFL2024 .
Join us and other AI & ML experts to find out the latest in ML for biology and chemistry. The call for talks is open!
Bruno Correia Philippe Schwaller (he/him) Arne Schneuing Ilia Igashov Rebecca Neeser
2024.appliedmldays.org/media-34-tracks


Join us at the AI & the Molecular World track in #AMLDEPFL2024 on 26 March 2024! 🧬
Call for talks: forms.gle/6GJ4GKvNQpv9Yb…
More info: 2024.appliedmldays.org/media-34-tracks


Excited to present Beam Enumeration at #ICLR2024 !
Check out the thread: Molecular substructures extracted from a generative model can be used for self-conditioned generation. The results are improved sample efficiency and providing explainability!
Thanks to my PI Philippe Schwaller (he/him)!

(Partially) made it to #NeurIPS2023 ! Check out our RetroBridge poster at NeurIPS AI4D3 Workshop (room 242) right now and at #AI4Science tomorrow 👀
We are not physically there but feel free to reach out here or by email!
Credits for pic to Rebecca Neeser


1/5 🌟 Thrilled to present our new work on predicting the synthesizability of small molecules this Saturday in the #AI4Science workshop at #NeurIPS2023 ! Thanks to my supervisors Philippe Schwaller (he/him) (@SchwallerGroup ) and Bruno Correia 🧵
schwallergroup.github.io/fsscore/FSscor…