Ruibang Laurent Luo
@aquaskyline
Associate Professor of Computer Science at HKU. Private Pilot. #bioinformatics/#engineering. Learning 🏌️ recently.
ID:109840044
http://luo-lab.hk 30-01-2010 13:10:52
788 Tweet
804 Takipçi
347 Takip Edilen

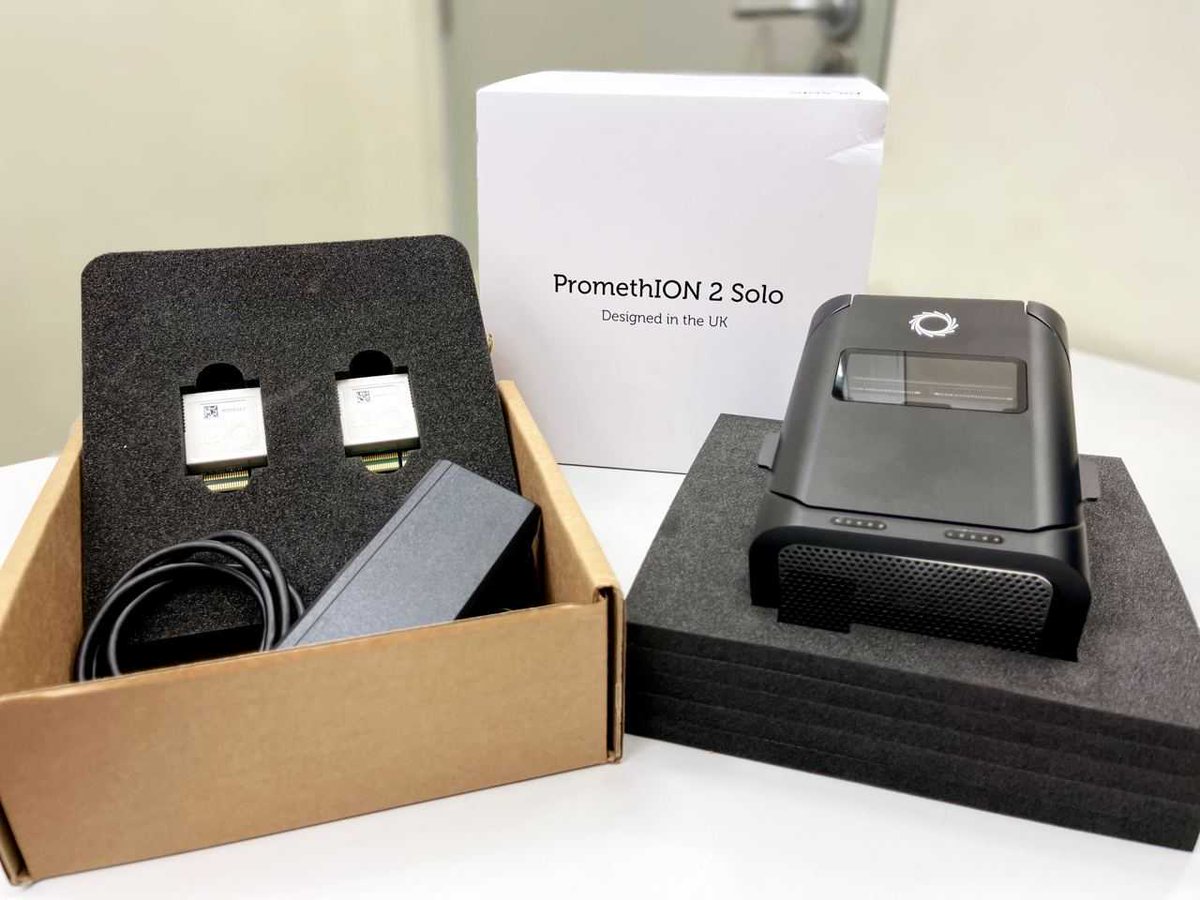
So excited to have received probably the first P2 solo in the greater China region today. Gonna use it for developing a somatic variant caller and many more exciting projects! Oxford Nanopore #precisionmedicine


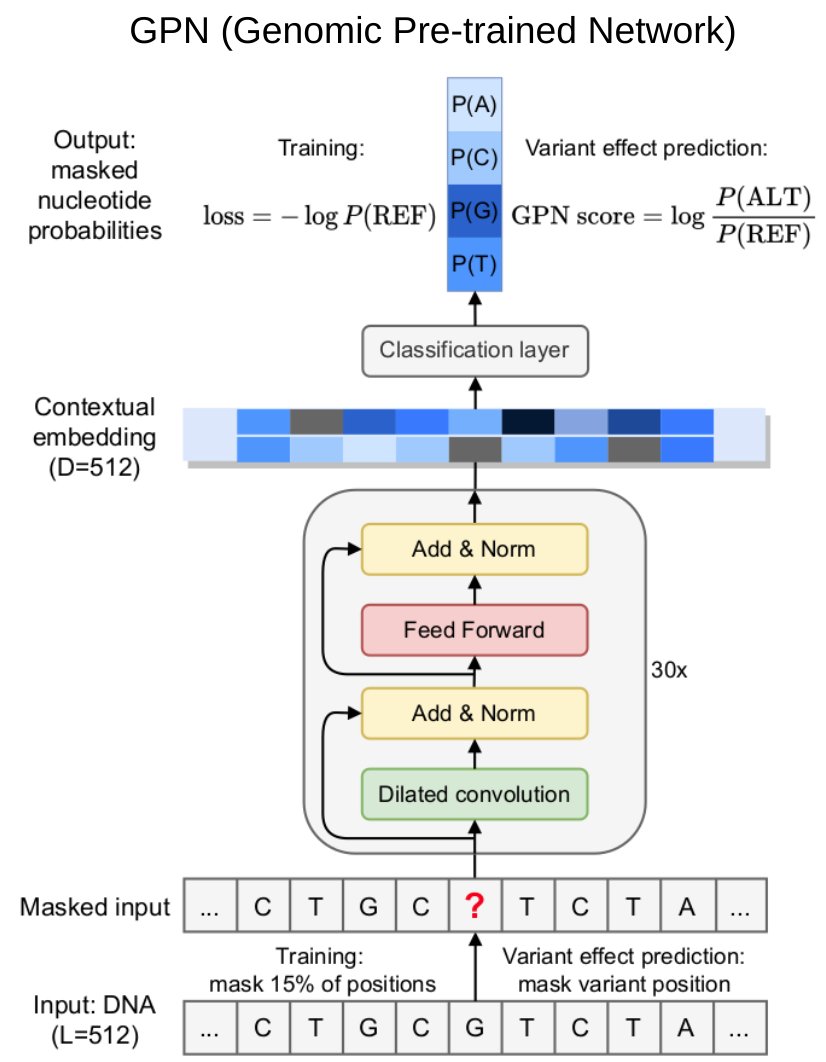
Excited to share our findings training GPN, a DNA language model for Arabidopsis thaliana, with @sanjitsbatra and Yun S. Song:
DNA language models are powerful predictors of non-coding variant effects, without the need for any labeled data.
doi.org/10.1101/2022.0…
1/n

A retrospective of our experience at #epi2melabs of using Nextflow to deliver bioinformatic workflows for Oxford Nanopore data analysis
labs.epi2me.io/two-years-of-n…



Please RT: We are looking for talented #Bioinformatics #postdocjobs who want to work on Oxford Nanopore & PacBio data from human population to diseases ( #Cancer , #Parkinsons , ..). Develop your own research and methods at scale. Apply: jobs.bcm.edu/job/Houston-Po… BCM HGSC From the Labs at Baylor College of Medicine


We have started collecting and uploading #RECOMBSeq2022 presentation videos to our official YouTube channel. More will be added as we receive them. The playlist is here: youtube.com/playlist?list=…



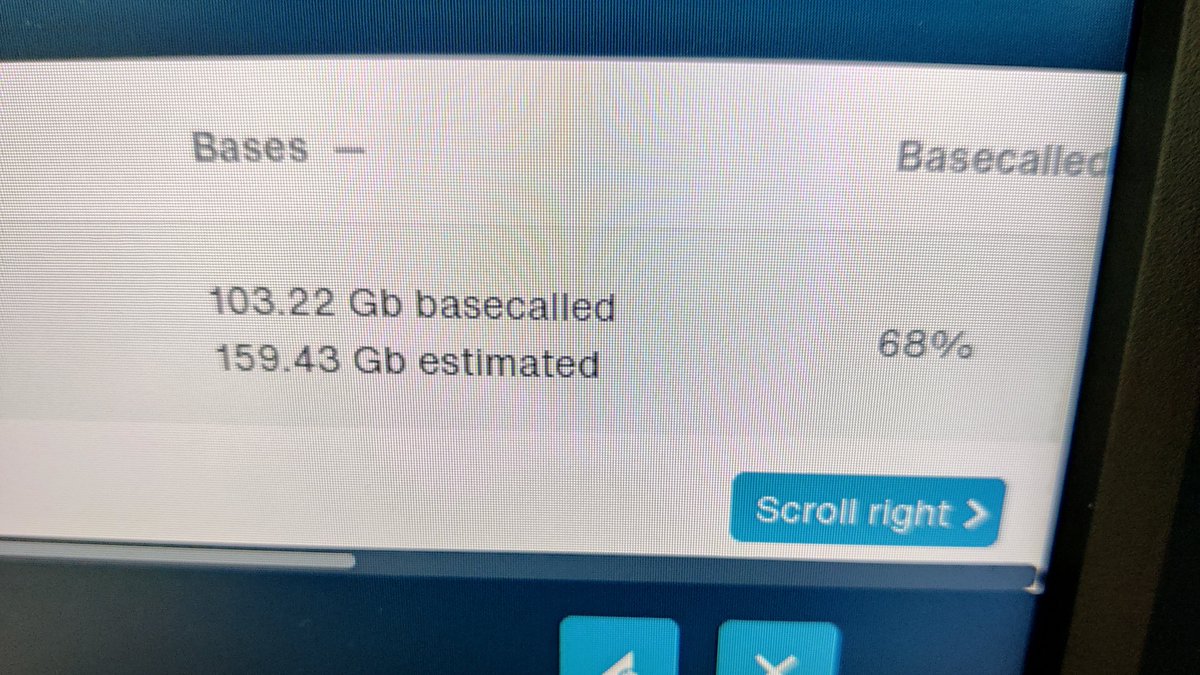
Kit14 on the Oxford Nanopore #PromethION bringing back kit10 like yields with kit12 like accuracy and sensitivity. Need a bigger GPU 🤣 60 ng of DNA loaded onto the flowcell.

The exciting reveal of Ultima Genomics last week was accompanied by the publication of four preprints. Intrigued by the potential of the technology, Sina Booeshaghi & I decided to take a look at the data. A 🧵 about our findings & a preprint we posted: biorxiv.org/content/10.110… 1/


After many months, I'm proud to share our work on untangling genome assembly graphs with GNNs. Or, GNNome Assembly (sorry not sorry)🧬
Paper: arxiv.org/abs/2206.00668
Code/Data: github.com/lvrcek/GNNome-…
with Xavier Bresson, T. Laurent, Martin Schmitz, and Mile Sikic.
Read more in 🧵
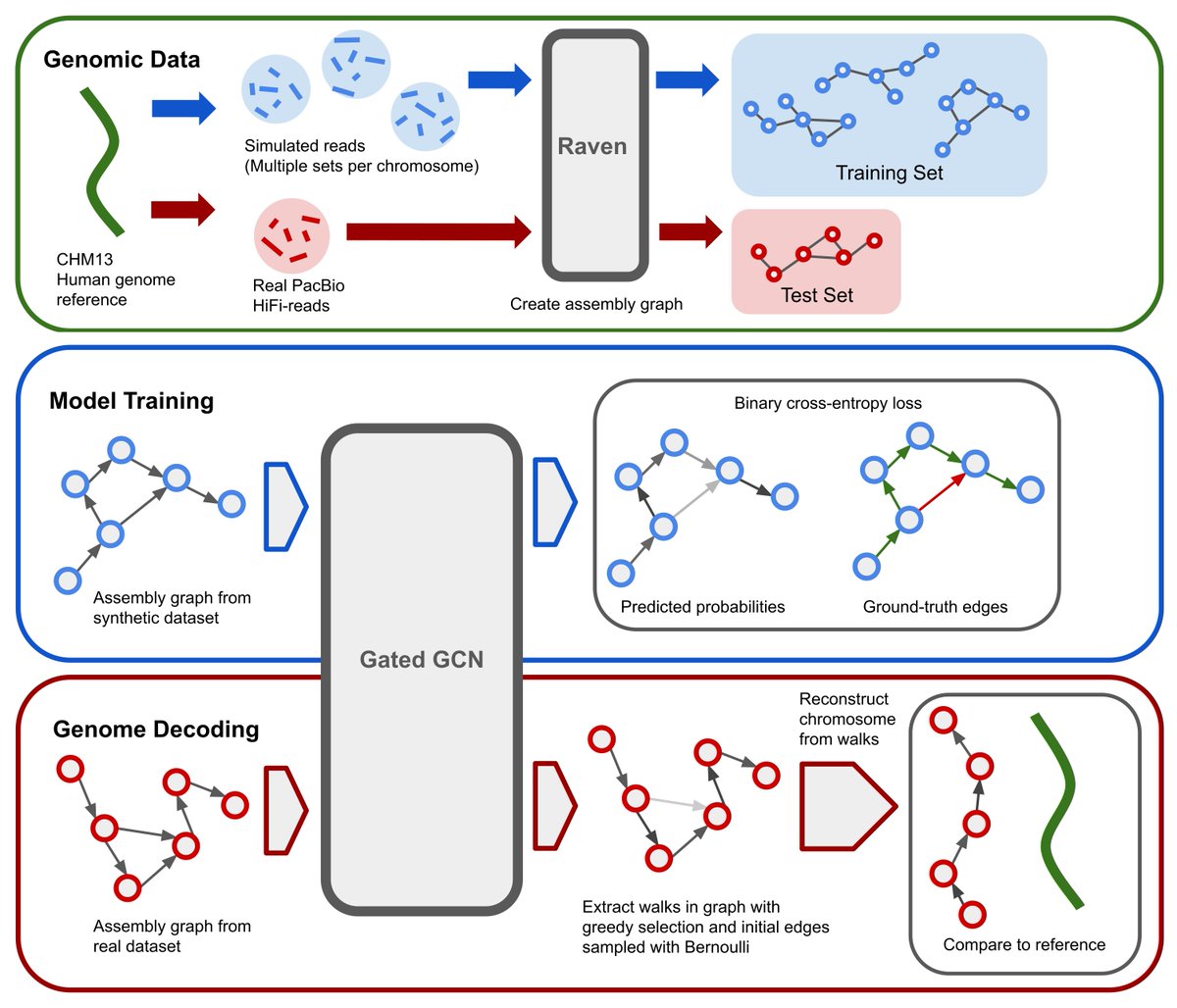

Image fraud gets a boost from AI
* Deepfakes in the biomedical literature are coming, if they aren't here already.
* GAN technology can easily create fake photos of Western blots or cancer cells.
Bradley van Paridon reports for Advanced Sci News
advancedsciencenews.com/image-fraud-ge…

hello #BoG22 attendees, I'm recruiting postdocs. See a list of former lab members at salzberg-lab.org/lab-members/ incl @carlk Martin Steinegger 🇺🇦 Cole Trapnell Michael Schatz Ben Langmead Daehwan Kim Ruibang Luo and others. If you want to join this group someday, email me!